SSGCID
Seattle Structural Genomics Center for Infectious Disease
SSGCID structures
Filter list by:
Results: 637
| Structure | PDB | Target | Annotation | Organism | Community | Deposited | Structure type | Ligands |
|---|---|---|---|---|---|---|---|---|
 |
7K62 | MykaA.01062.a | Dihydrofolate reductase (EC 1.5.1.3) | Mycobacterium kansasii | request | 09/18/2020 | XRay | NADP;p218; |
 |
6Q09 | MykaA.20209.a | Hemerythrin HHE cation binding domain-containing protein; Rv2633c homolog | Mycobacterium kansasii | request | 08/01/2019 | XRay | iron; |
 |
6TYJ | MykaA.20209.a | Hemerythrin HHE cation binding domain-containing protein; Rv2633c homolog | Mycobacterium kansasii | request | 08/09/2019 | XRay | zinc; |
 |
6U3L | MykaA.20209.a | Hemerythrin HHE cation binding domain-containing protein; Rv2633c homolog | Mycobacterium kansasii | request | 08/22/2019 | XRay | — |
 |
4EO9 | MyleA.00184.a | PROBABLE PHOSPHOGLYCERATE MUTASE 1 GPM1 (PHOSPHOGLYCEROMUTASE) (PGAM) (BPG-DEPENDENT PGAM) | Mycobacterium leprae | request | 04/13/2012 | XRay | — |
 |
4J07 | MyleA.00730.a | PROBABLE RIBOFLAVIN SYNTHASE BETA CHAIN RIBH (6,7-dimethyl-8-ribityllumazine synthase) (DMRL synthase) (Lumazine synthase) | Mycobacterium leprae | request | 01/30/2013 | XRay | — |
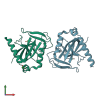 |
4ECP | MyleA.01382.a | INORGANIC PYROPHOSPHATASE PPA (PYROPHOSPHATE PHOSPHO-HYDROLASE) (PPASE) (INORGANIC DIPHOSPHATASE) (DIPHOSPHATE PHOSPHO-HYDROLASE | Mycobacterium leprae | request | 03/26/2012 | XRay | — |
 |
4WKW | MyleA.18251.a | CONSERVED HYPOTHETICAL PROTEIN | Mycobacterium leprae | request | 10/03/2014 | XRay | — |
 |
4EX4 | MyleA.18344.a | PROBABLE MALATE SYNTHASE G GLCB | Mycobacterium leprae | request | 04/29/2012 | XRay | — |
 |
6D47 | MymaA.17987.a | DNA polymerase III subunit beta (EC 2.7.7.7) | Mycobacterium marinum | request | 04/17/2018 | XRay | — |
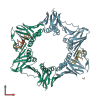 |
6DLY | MymaA.17987.a | DNA polymerase III subunit beta (EC 2.7.7.7) | Mycobacterium marinum | request | 06/04/2018 | XRay | Griselimycin; |
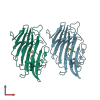 |
4PQ9 | MymaA.18623.b | Beta-1,3-glucanase (Rv0315 ortholog) | Mycobacterium marinum | request | 02/28/2014 | XRay | — |
 |
6Q10 | MymaA.19257.a | Conserved hypothetical secreted protein (Rv2700 ortholog) | Mycobacterium marinum | request | 08/02/2019 | XRay | — |
 |
4XXP | MypaA.18623.a | Putative uncharacterized protein (Rv0315 ortholog) | Mycobacterium paratuberculosis | request | 01/30/2015 | XRay | magnesium; |
 |
6N39 | MypaA.19484.a | Dephospho-CoA kinase CoaE | Mycobacterium paratuberculosis | request | 11/14/2018 | XRay | — |
 |
5VO7 | MysmA.01550.a | Thioredoxin (Rv1471 ortholog) | Mycobacterium smegmatis | request | 05/02/2017 | NMR | — |
 |
4PFS | MysmA.11416.b | Cobyrinic Acid a,c-diamide synthase (Rv3918c ortholog) | Mycobacterium smegmatis | request | 04/30/2014 | XRay | — |
 |
5IF9 | MysmA.11416.b | Cobyrinic Acid a,c-diamide synthase (Rv3918c ortholog) | Mycobacterium smegmatis | request | 02/25/2016 | XRay | magnesium;AMP-PNP; |
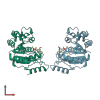 |
5IHP | MysmA.11416.b | Cobyrinic Acid a,c-diamide synthase (Rv3918c ortholog) | Mycobacterium smegmatis | request | 02/29/2016 | XRay | ADP;magnesium; |
 |
6PI4 | MysmA.18337.a | ATP synthase epsilon chain (ATP synthase F1 sector epsilon subunit) (F-ATPase epsilon subunit) | Mycobacterium smegmatis | request | 06/25/2019 | XRay | — |
 |
4TMD | MysmA.18640.a | Putative uncharacterized protein (Rv0999 ortholog) | Mycobacterium smegmatis | request | 06/01/2014 | XRay | — |
 |
6AQH | MythA.00612.a | Lysine--tRNA ligase LysS (EC 6.1.1.6) (Lysyl-tRNA synthetase) | Mycobacterium thermoresistibile | request | 08/19/2017 | XRay | L-Lysine;Cladosporin; |
 |
4U7J | MythA.00809.a | Argininosuccinate synthase (EC 6.3.4.5) (Citrulline--aspartate ligase) (Rv1658 ortholog) | Mycobacterium thermoresistibile | request | 07/30/2014 | XRay | — |
 |
4XFJ | MythA.00809.a | Argininosuccinate synthase (EC 6.3.4.5) (Citrulline--aspartate ligase) (Rv1658 ortholog) | Mycobacterium thermoresistibile | request | 12/27/2014 | XRay | AMP-PNP; |
 |
6AP5 | MythA.00916.a | Thioredoxin (Rv1471 ortholog) | Mycobacterium thermoresistibile | request | 08/17/2017 | NMR | — |