Mechanisms of persistent bacterial infection
Several pathogenic bacteria are able to survive inside their host for extended periods of time, causing recurrent disease. The group of Dr. Christoph Dehio, from the Biozentrum Basel, Switzerland, has established a long-standing partnership with the SSGCID to uncover the mechanisms of persistent bacterial infection, which are still poorly understood, with the ultimate goal of designing new drugs to combat chronic infectious disease. In particular, the collaboration with SSGCID focuses on characterizing the structure-function relationships of Bartonella effector proteins (Beps), which play a critical role in persistent infection.
Bartonella, a gram-negative bacterium, is a choice model for studying persistent infection: it is able to survive in numerous animal reservoir hosts. In order to establish persistent infection, Bartonella injects a cocktail of effector proteins through the type IV secretion system (T4SS) into host cells. Once injected, the effector proteins alter multiple signaling pathways to enable survival of the bacteria inside the host. From their colonized niche, the bacteria are then periodically released into the bloodstream, causing the typical relapsing Bartonella infections (PMID: 22229763).
The terminal BID domain of the Beps is not only essential for successful injection of effector proteins, but it also aids in establishing persistent infection by interacting with host proteins (PMID: 25461577). The Dehio group studies this ability to perform multiple functions using sequence analysis, biochemistry and cell biology techniques. They demonstrated that BID domains belong to different classes, each with distinct secondary functions (PMID: 21347280). BID domains were also found in a family of relaxases, proteins responsible for the transfer of plasmid and chromosomal DNA from one bacterium to another during conjugation (PMID: 23995644). However, in the absence of structural information, the molecular basis for the functional diversity of the BID domain remains an open question.
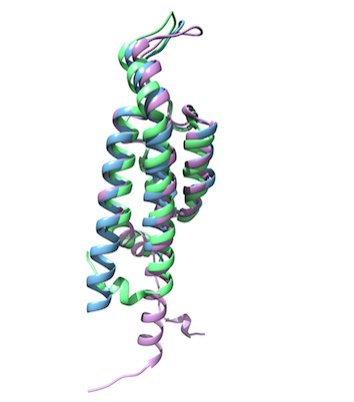
The SSGCID successfully determined the first structures of BID domains from 3 different classes. Detailed analysis of the structures revealed that the dual function of BID domains is reflected in their structure, which comprises a conserved, novel, rigid fold for secretion and a variable surface to facilitate the evolution of secondary functions inside host cells. The compact and elongated shape of the BID domain is compatible with translocation via the T4SS in a fully folded state, while the presence of an invariantly positively charged “hook” suggests a conserved translocation mechanism. This work is a stepping stone towards understanding, at the molecular level, how stealth bacterial pathogens such as Bartonella inject virulence factors into their hosts and establish persistent infection. Efforts are underway to confirm the role of the “hook” by site-directed mutagenesis and to investigate interactions with host partners using various methods, including chemical cross-linking and mass spectrometry.
Details of this work have been published in the journal Structure. Reference: The BID domain of Type IV Secretion Substrates forms a conserved four-helix bundle topped with a hook. Stanger FV, de Beer TA, Dranow DM, Schirmer T, Phan I, Dehio C. Structure Epub ahead of print (Pubmed ID 27889208). Coordinates are available in the Protein Data Bank www.rcsb.org, PDB IDs 4YK1, 4YK2, 4YK3.